linux上安装methylkit -- 安全下车版 (正经版: Linux环境下安装methylKit的实践与避坑指南)
题外话:
我踩过的坑,都将成为我写贴的素材!(ㄒoㄒ)
整整安装了两天,这里面的滋味懂的都懂。
希望开发作者持续维护。
希望有人或者作者持续打包成sigularity镜像使用,并且直接传到github上,传到docker上下载到linux上也好困难啊(ㄒoㄒ) ,这样就不会吃conda安装的苦了。
核心思想:
- 用mamba 或者conda 安装,这样依赖包都同时安装好了。
- 网要好。我的实践是早上网最好,比如7-9点时,我的下载网速最高能达到1.2M/s (因为就算设了清华源等国内源,还是有一部分需要链接到国外网址的,这个时候网速就十分受限。所以需要与国外上班族错峰使用。)
安装过程中出现这个报错就是网不好造成的。👉conda或mamba install 相关软件报错 - 安装data.table版本为1.14.8
安装过程中报错
Error: package or namespace load failed for 'methylKit':object 'key<-' is not exported by 'namespace:data.table'
详细解释看后续部分内容。
解决办法就是安装data.table到1.14.8或更低版本。👉2025.06.23【甲基化】|methylKit常见疑难解答与实用技巧(FAQ)
但是博主的方法我用了不成功,所以这里提供另一种方法。
正式开始:
- 创建环境并安装所需包,部分细节看👉被迫在linux上用R(真的很难用啊)之如何在linux上正常使用R
mamba create -n methylkit_5 -c bioconda -c conda-forge bioconductor-methylkit bioconductor-genomation r-data.table=1.14.8 -y
bioconductor-methylkit 必须安装
bioconductor-genomation 是为了后续将差异甲基化位点注释到基因组上
r-data.table=1.14.8 必须指定版本安装,让mamba自动匹配合适的methylkit和genomation。(这就是我说的另一种方法,从安装的时候就指定版本,这是我折腾最久的地方,不多说了。)
安装关键过程:
+ bioconductor-methylkit 1.28.0 r43hf17093f_1 bioconda Cached
+ bioconductor-genomation 1.34.0 r43hf17093f_1 bioconda Cached
+ r-base 4.3.3 h65010dc_18 conda-forge Cached
+ r-data.table 1.14.8 r43h029312a_2 conda-forge Cache...
Downloading and Extracting Packages:Preparing transaction: done
Verifying transaction: done
Executing transaction: doneTo activate this environment, use$ mamba activate methylkit_5To deactivate an active environment, use$ mamba deactivate这里不禁要提一下原作者github上的issue部分:
原链接👉添加链接描述
防止有人打不开,这里再附一张截图
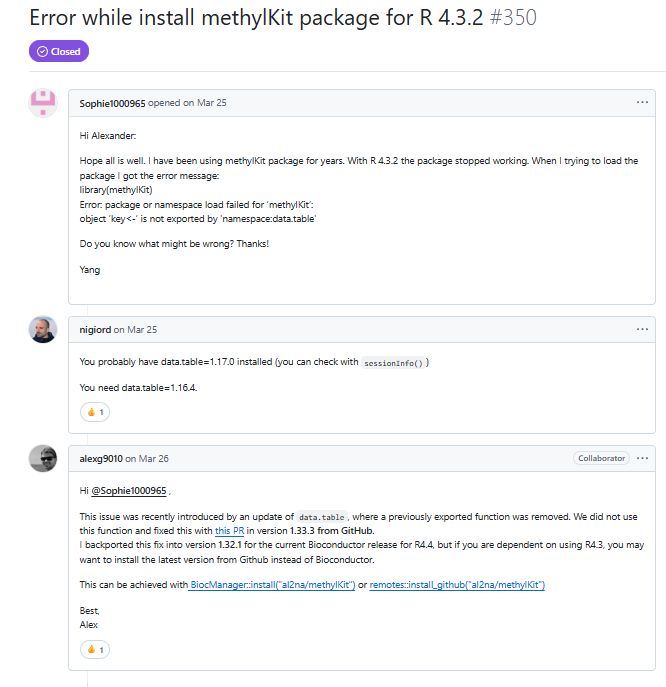
这里作者解释了,这个报错就是因为data.table更新了,新版本删除了一个函数。作者修复了这一报错,并建议从R版本为4.4的Bioconductor下载1.32.1版本的methylKit。(原文自己细看,这里是我的理解。)
但是bioconda里的版本截至到目前只更新到1.32.0(2025/8/29),这意味着linux上安装methylkit建议还是安装data.table的旧版本(1.14.8),methylkit安装的版本也是旧版本,比如1.28.0。同时,也意味着,如果在电脑本地安装,可以将R升级到4.4版本,然后Bioconductor就可以安装methylkit的最新版本了。
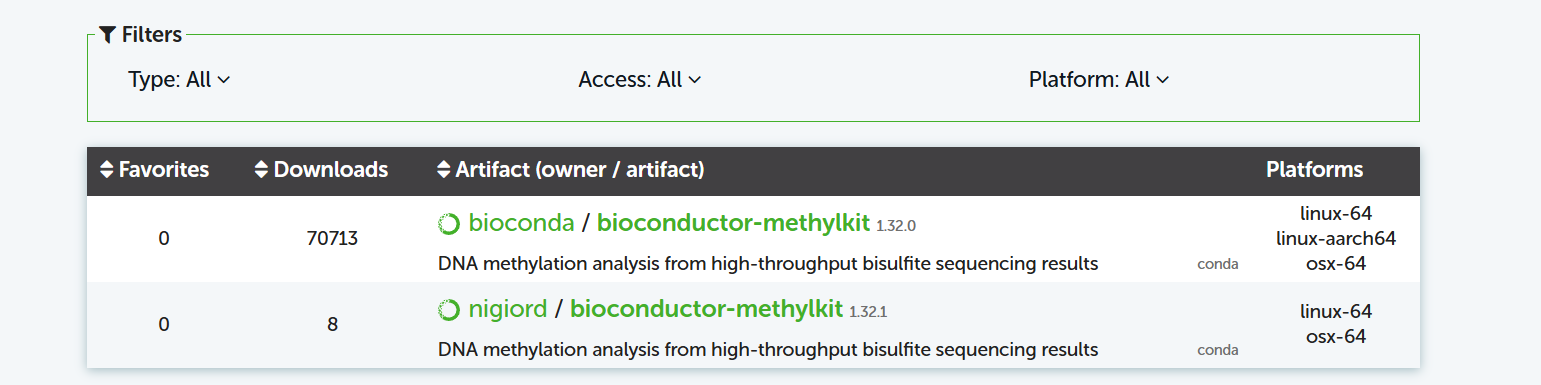
(nigiord的源可能不靠谱,不建议尝试,头铁也可以试试安装看下有没有问题。)
- 检测是否安装成功
① 激活环境
mamba activate methylkit_5
② 打开R
直接命令行中输入R,进入R
输入.libPaths() #查看当前R包路径
临时改成该环境methylkit_5的R包路径
.libPaths(c("/storage2/zuozhe/mambaforge/envs/methylkit_5/lib/R/library", .libPaths()))
这里每个人的路径都不一样,如果不确定可以去翻mambaforge/envs/methylkit_5/lib/R/library
就能得到自己的绝对路径了。
③ library对应包,如果没出现报错说明就安装成功了,正常使用。
(methylkit_5) zuozhe@server:~$ RR version 4.3.3 (2024-02-29) -- "Angel Food Cake"
Copyright (C) 2024 The R Foundation for Statistical Computing
Platform: x86_64-conda-linux-gnu (64-bit)R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> .libPaths()
> .libPaths(c("/storage2/zuozhe/mambaforge/envs/methylkit_5/lib/R/library", .libPaths()))
> library(methylKit)
Loading required package: GenomicRanges
Loading required package: stats4
Loading required package: BiocGenericsAttaching package: 'BiocGenerics'The following objects are masked from 'package:dplyr':combine, intersect, setdiff, unionThe following objects are masked from 'package:stats':IQR, mad, sd, var, xtabsThe following objects are masked from 'package:base':Filter, Find, Map, Position, Reduce, anyDuplicated, aperm, append,as.data.frame, basename, cbind, colnames, dirname, do.call,duplicated, eval, evalq, get, grep, grepl, intersect, is.unsorted,lapply, mapply, match, mget, order, paste, pmax, pmax.int, pmin,pmin.int, rank, rbind, rownames, sapply, setdiff, sort, table,tapply, union, unique, unsplit, which.max, which.minLoading required package: S4VectorsAttaching package: 'S4Vectors'The following objects are masked from 'package:dplyr':first, renameThe following object is masked from 'package:utils':findMatchesThe following objects are masked from 'package:base':I, expand.grid, unnameLoading required package: IRangesAttaching package: 'IRanges'The following objects are masked from 'package:dplyr':collapse, desc, sliceLoading required package: GenomeInfoDbAttaching package: 'methylKit'The following object is masked from 'package:dplyr':select> library(genomation)
Loading required package: gridAttaching package: 'genomation'The following objects are masked from 'package:methylKit':getFeatsWithTargetsStats, getFlanks, getMembers,getTargetAnnotationStats, plotTargetAnnotationWarning message:
replacing previous import 'Biostrings::pattern' by 'grid::pattern' when loading 'genomation'④ 如果library过程或者运行methylkit中报错了缺啥包,就缺啥包再安装啥包 (Warning message 影响不大)。这种情况没什么更好的经验,可以尝试在R中直接安装,也可以在bioconda中找到对应的包安装,再重新library。如果折腾来折腾去都没成功,建议重新建个环境,加上所需的包,比如我的就要求加上dplyr包,所以最终我的命令是:
mamba create -n methylkit_5 -c bioconda -c conda-forge bioconductor-methylkit bioconductor-genomation r-data.table=1.14.8 r-dplyr -y
目前就是这样,研究methylkit命令去了。没看到最后不知道还有个最终命令吧。
